
Novel carbohydrate sulfotransferase 3 mutation causing spondyloepiphyseal dysplasia with congenital joint dislocations in a Chinese family
Spondyloepiphyseal dysplasia with congenital joint dislocations (SEDCJD; OMIM 143095) is an autosomal recessive skeletal dysplasia with clinic features such as severe progressive kyphoscoliosis, arthritic changes with joint dislocations, genu valgum, cubitus valgus, mild brachydactyly, camptodactyly, microdontia, and normal intelligence. The affected individuals may receive an initial clinical diagnosis of Larsen syndrome (OMIM 245600) or humerospinal dysostosis (1). SEDCJD is usually evident at birth. The dislocations were aggravated year by year, and the features of spondyloepiphyseal dysplasia become apparent during grow-up. These will lead to arthritis changes of the hip and spine, rigid kyphoscoliosis, and trunk shortening (2). Congenital valvular heart disease has also been reported in several patients with SEDCJD (2-6). Our proband had heart problem with mitral regurgitation, and has had a son, who died of congenital heart disease at the age of 7 months. It is likely that involvement of cardiac valvular connective tissue is an important feature of SEDCJD.
SEDCJD was caused by carbohydrate sulfotransferase 3 (CHST3, OMIM 603799) mutations. CHST3 spans more than 20 kb with three exons, encodes human C6ST, which consist of a core protein that distributed on the surfaces of most cells and the extracellular matrix in virtually every tissue (4). In this study, we presented the clinical manifestations and radiographic features of one Chinese SEDCJD patient, whom we found a novel CHST3 missense mutation.
The 35-year-old male proband is the only child born to healthy but consanguineous parents (cousins) of Chinese Han descent. There was no family history of SEDCJD or other skeletal diseases. The hip joint dislocations and elbow subluxations of the proband were observed at birth. At the age of 10, he began to suffer from severe hip joint pain and had to rely on wheelchair. At the age of 15, his stature appeared to be significantly shorter than his peers. Short trunk with intervertebral disc degeneration and rigid thoracic kyphoscoliosis were noted. The proband has a rigid lumbar spine with flexibility less than 30%. At the age of 19, the proband underwent treatment of the femoral head osteonecrosis by using bone flap with blood pedicle and cannulated screw internal fixation. Progression of knee epiphyseal changes, osteopenia, short and broad thumbs (Figure 1) and a lordotic gait were also present at this time. Neurological and other examinations were within the normal range. His height was 145 cm (−3.8 SD).
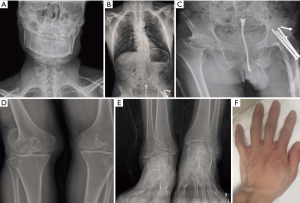
X-ray imaging at 35 years revealed a high palate and short neck (Figure 1A). A fusion of vertebral bodies, severe scoliosis and narrowed intervertebral space were noted (Figure 1B). There were significant bilateral hip joint destruction and generalized osteopenia. Short femoral neck and severe osteoarthritis observed (Figure 1C). The femoral epiphyses were enlarged, irregular and small epiphyses of the tibia were noted (Figure 1D). Clubfoot was discovered and the joint space was narrow (Figure 1E).
In retrospect, the radiographic findings of the proband were quite identified with the previous reports of SEDCJD (6-8); however, because of the closure of growth plates and secondary changes related to the aging, definite diagnosis was difficult. Therefore, we performed molecular analysis of the proband. After the written informed consent, genomic DNAs from peripheral blood of the proband and his family was extracted using a QIA-GEN DNA minikit (Qiagen, Hilden, Germany).
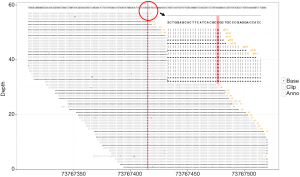
The whole exome sequencing (WES) in the proband was conducted. Briefly, genomic DNA was divided into smaller fragments of 200–250 bp by ultrasonic instrument (Covaris LE220, Massachusetts, USA). Subsequently, Ampure Beads (Beckman Coulter, California, USA) purification was used to add poly A/joint reaction in the end of the purified DNA fragmentS The gene-trapping chip (Roche NimbleGen, Madison, USA) was used to hybridize with the purified DNA fragments of proband. After hybridization, captured DNAs was sequenced on Illumina HiSeq2500 Analyzers (Illumina, SanDiego, USA) and read on Illumina Pipeline software (version 1.3.4). We used BWA v0.59 (9) to align sequence reads to the human genome reference (build 37) and removed duplicated reads from subsequent analyses. Sequences variants were identified by compared to the NCBI reference sequence (NM 004273.4) and annotated by ANNOVAR (http://www.openbioinformatics.org/annovar). The 20× coverage for the RefSeq coding region was 97.47% (Figure S1).
One novel missense mutations (c.626 C>A; p.Pro209Gln) in exon 3 of CHST3 (Figure 2A) was discovered in the proband. This mutation was not previously reported and was not found in sequencing data projects (Exome Aggregation Consortium Exome http://exac.broadinstitute.org/). One cytosine-ribonucleotide altered to adenine-ribonucleotide in codon 626 and caused the change of reading frame from proline to glutamine. SIFT and Polyphen-2 (10) both predicted the mutations to have severe damage to the CHST3 protein function.
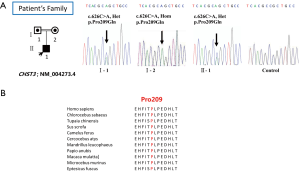
We performed Sanger sequencing for the proband’s family and confirmed the mutation c.626C>A; p.Pro209Gln (Figure 2A) which was highly conserved among diverse species (Figure 2B). The proband’s parents are healthy, and they are carrier of this missense mutation. All this suggested that mutation c.626C>A was pathopoiesis. The inbreeding caused the proband’s deformity, a homozygote missense mutation of CHST3.
Thus, one novel CHST3 mutation in an adult case of SEDCJD was discovered in this study. Our study performed the WES, which allows us to get molecular results faster especially when traditional specific diagnosis takes a long time to sequence a large gene. The exome will give us possibility to analyze the first candidate gene and check the presence of mutations in other genes at the same time (11,12). This study has further expanded the CHST3 mutation spectrum, and thus contributed to the early detection of this disease, providing significant benefits to the patient and his families, and being aware of the growing amount of SEDCJD patients for future clinic diagnosis.
Acknowledgments
The authors thank the patients and their families who donated their blood samples for this study.
Funding: This work was supported by the Projects of International Cooperation and Exchanges NSFC (81420108021), National Key Technology Support Program (2015BAI08B02), Excellent Young Scholars NSFC (81622033), Jiangsu Provincial Key Medical Center Foundation, Jiangsu Provincial Medical Talent Foundation and Jiangsu Provincial Medical Outstanding Talent Foundation.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/aoj.2017.01.02). QJ serves as an Editor-in-Chief of Annals of Joint from Mar 2016 to Feb 2021. DS serves as an unpaid Executive Editor-in-Chief of Annals of Joint from Mar 2016 to Feb 2021. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee(s) and with the Helsinki Declaration (as revised in 2013). Written informed consent was obtained from the patient for publication of this manuscript and any accompanying images.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Bayoumi R, Saar K, Lee YA, et al. Localisation of a gene for an autosomal recessive syndrome of macrocephaly, multiple epiphyseal dysplasia, and distinctive facies to chromosome 15q26. J Med Genet 2001;38:369-73. [Crossref] [PubMed]
- Unger S, Lausch E, Rossi A, et al. Phenotypic features of carbohydrate sulfotransferase 3 (CHST3) deficiency in 24 patients: congenital dislocations and vertebral changes as principal diagnostic features. Am J Med Genet A 2010;152A:2543-9. [Crossref] [PubMed]
- Tanteles GA, Dixit A, Dhar S, et al. Two Somali half-siblings with CHST3-related chondrodysplasia illustrating the phenotypic spectrum and intrafamilial variability. Am J Med Genet A 2013;161A:2588-93. [PubMed]
- van Roij MH, Mizumoto S, Yamada S, et al. Spondyloepiphyseal dysplasia, Omani type: further definition of the phenotype. Am J Med Genet A 2008;146A:2376-84. [Crossref] [PubMed]
- Hermanns P, Unger S, Rossi A, et al. Congenital joint dislocations caused by carbohydrate sulfotransferase 3 deficiency in recessive Larsen syndrome and humero-spinal dysostosis. Am J Hum Genet 2008;82:1368-74. [Crossref] [PubMed]
- Thiele H, Sakano M, Kitagawa H, et al. Loss of chondroitin 6-O-sulfotransferase-1 function results in severe human chondrodysplasia with progressive spinal involvement. Proc Natl Acad Sci U S A 2004;101:10155-60. [Crossref] [PubMed]
- Tuysuz B, Mizumoto S, Sugahara K, et al. Omani-type spondyloepiphyseal dysplasia with cardiac involvement caused by a missense mutation in CHST3. Clin Genet 2009;75:375-83. [Crossref] [PubMed]
- Rajab A, Kunze J, Mundlos S. Spondyloepiphyseal dysplasia Omani type: a new recessive type of SED with progressive spinal involvement. Am J Med Genet A 2004;126A:413-9. [Crossref] [PubMed]
- Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010;26:589-95. [Crossref] [PubMed]
- Kelley LA, Sternberg MJ. Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 2009;4:363-71. [Crossref] [PubMed]
- Lim BC, Yoo SK, Lee S, et al. Hoyeraal-Hreidarsson syndrome with a DKC1 mutation identified by whole-exome sequencing. Gene 2014;546:425-9. [Crossref] [PubMed]
- Mansouri M, Kayserili H, Elalaoui SC, et al. Novel DDR2 mutation identified by whole exome sequencing in a Moroccan patient with spondylo-meta-epiphyseal dysplasia, short limb-abnormal calcification type. Am J Med Genet A 2016;170A:460-5. [Crossref] [PubMed]
Cite this article as: Yan W, Dai J, Shi D, Xu X, Chen D, Xu Z, Song K, Yao Y, Teng H, Jiang Q. Novel carbohydrate sulfotransferase 3 mutation causing spondyloepiphyseal dysplasia with congenital joint dislocations in a Chinese family. Ann Joint 2017;2:7.


